|
5/30/2023 0 Comments Enzymex primer verification 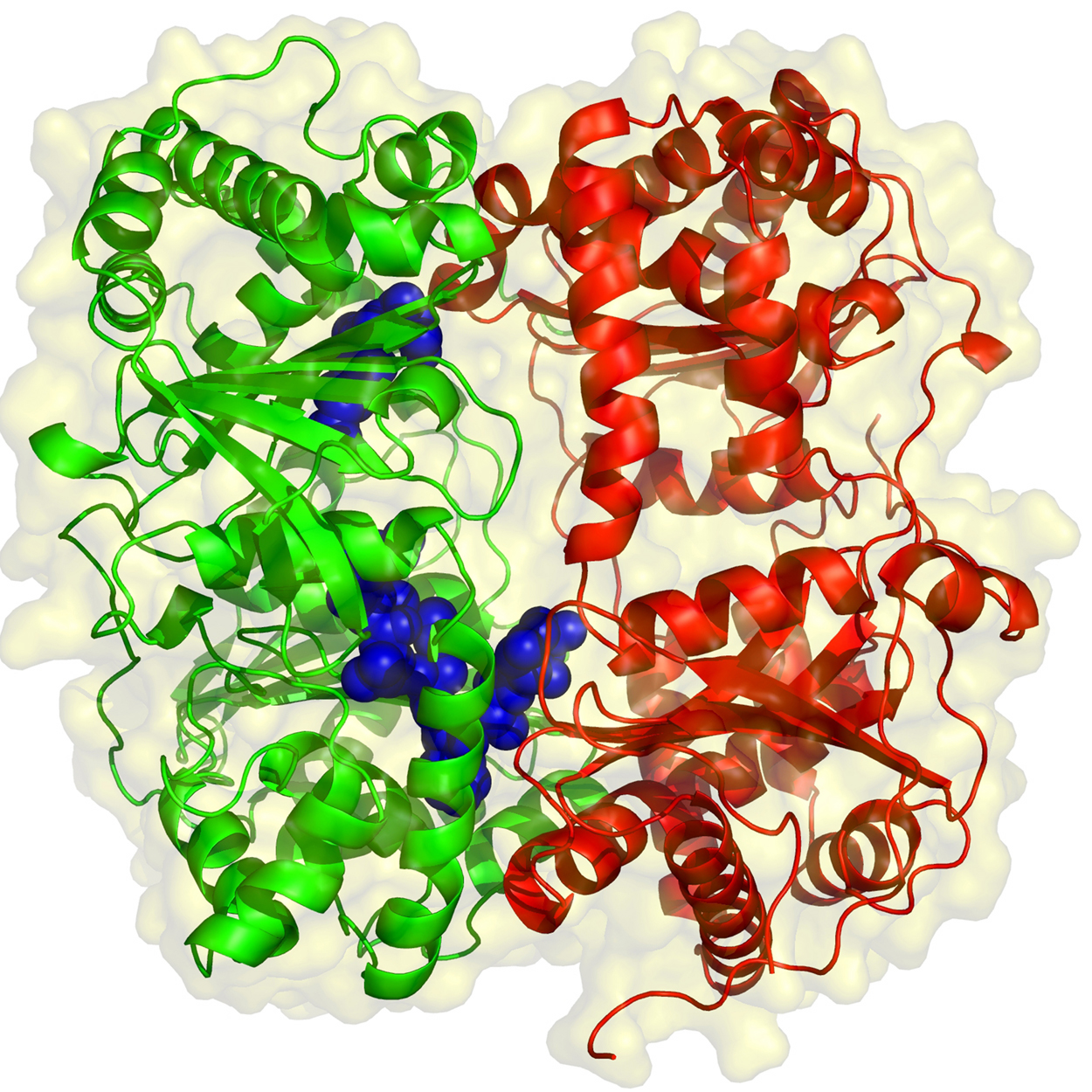
In the diagram, the primosome is shown moving in a 5' to 3' direction along the template strand (clockwise), which is the opposite of the direction of primer synthesis. This tracking along the SSB-coated single stranded DNA requires ATP hydrolysis and causes dissociation of some of the SSB. After synthesis of the primer (dark black line), the primosome moves to the next site for synthesis. 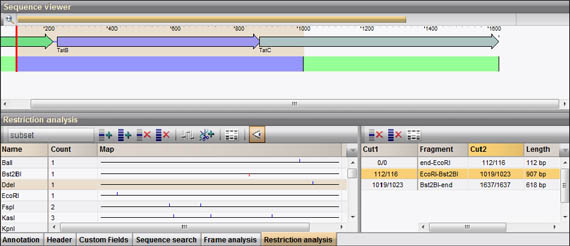
After assembly of the prepriming complex, DnaG joins the complex to complete the primosome. The primosome can now move to the next site for primer synthesis. After ATP hydrolysis by DnaB, the low affinity ADP-DnaB complex dissociates from the template. A high affinity complex between DnaB and ATP forms or stabilizes a secondary structure in the single-stranded template DNA that is used by primase this is thought to be how DnaB "activates" the primase to begin synthesis. The preferred binding site on the template for primase is CTG, with the T being used as the template for the first nucleotide of the primers. Although the role of each of the proteins in the primosome is not yet clear, information is available on some of the steps in primosome action. The prepriming complex is now ready for the primase, DnaG, to bind and make the active primosome. In an ATP-dependent process, and with help from DnaT, DnaB is transferred to the template and DnaC is released. The hexameric protein DnaBis in a complex with six molecules of DnaCwhen it is not on the DNA. The proteins PriBand PriCare then added to form a complex. In the case of fX174 viral DNA template coated with SSB, PriA (Figure 5.24 B) recognizes a primer assembly site. A sixth protein, DnaC, is needed for the assembly of this complex. DNA dependent ATPase.įive different proteins are found in a prepriming complex, PriA, PriB, PriC, DnaT, and DnaB (Table 5.4). Needed to add DnaB‑DnaC complex to preprimosome Helicase, 3' to 5' movement, site recognition 
This reaction can be carried out in vitro, which allowed the biochemical dissection of the various steps in primosome assembly and movement. The conversion of single‑stranded phage DNA to duplex DNA occurs by the synthesis of several Okazaki fragments, and hence it is a good model for discontinuous synthesis on the lagging strand. coli, this plus strand is converted to a double-stranded replicative form (Figure 5.24 A). fX174 is a single-stranded bacteriophage the DNA found in the virus is termed the plus strand. Identification of the components of the primosome was aided by the convenient model system of in vitrosynthesis of fX174 DNA. The primosome acts repeatedly during lagging strand synthesis, finding a primer-binding site on the SSB-coated single-stranded template strand and synthesizing a primer. coli, DnaG protein, cannot synthesize primers by itself, but rather it is part of much larger complex called the primosome. The major primase in eukaryotic cells is DNA polymerase \(\alpha\). coli is the 60 kDa protein called DnaG protein, the product of the dnaG gene. They can be made of ribonucleotides or a mixture of deoxyribonucleotides and ribonucleotides. Primers are short oligonucleotides, ranging from 6 to 60 nucleotides long. The enzyme primase catalyzes the synthesis of the primers from which DNA polymerases can begin synthesis (Figure 5.21). We also strongly urge researchers to report as much information about their assays as possible in their publications.\) We present an overview of the main steps in the primer design workflow, with data that illustrate some of the unexpected variability that often occurs when theory is translated into practice. Despite the framework provided by the MIQE guidelines and the accessibility of wide-ranging support from peer-reviewed publications, books and online sources as well as commercial companies, the design of many published assays continues to be less than optimal: primers often lack intended specificity, can form dimers, compete with template secondary structures at the primer binding sites or hybridise only within a narrow temperature range. Consequently, poor design combined with failure to optimise reaction conditions is likely to result in reduced technical precision and false positive or negative detection of amplification targets. Primers are arguably the single most critical components of any PCR assay, as their properties control the exquisite specificity and sensitivity that make this method uniquely powerful. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
